Heart disease is a leading cause of death worldwide. Overall, 697,000 people in the United States die from heart disease, about one in every five deaths. The most common type is Coronary Heart Disease (CHD), a condition in which the arteries cannot deliver enough oxygen-rich blood to the heart. Did you know that certain genetic variants may protect against heart disease?
Multiple factors affect heart health, including age, genetics, other health conditions, and lifestyle choices. Some factors cannot be controlled. For example, advanced age, being born male, and family history increases your risk.
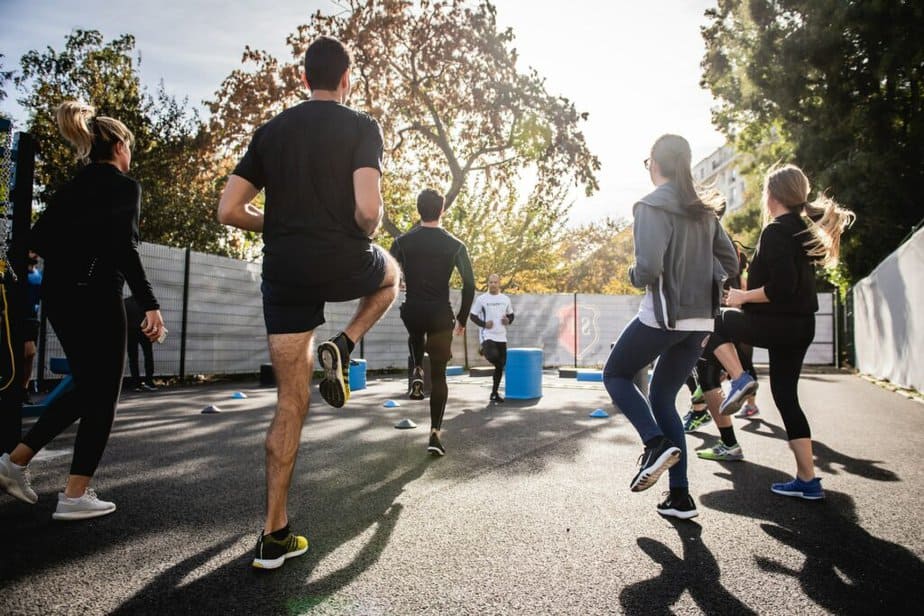
Patients can often mitigate other risk factors, such as smoking, lack of exercise, and obesity by lifestyle choices. Additionally, other diseases that affect heart health, such as diabetes and high cholesterol, can be managed with the help of your doctor.
| You can use the Nebula Genome Browser to search for these variants in your genome. Keep reading to learn what these newly discovered mutations are and how you can check your status. |
What causes CHD?
Several things can damage arteries, including diabetes, high blood pressure, high cholesterol, lack of exercise, and smoking. Patients can often prevent or manage most of these risks. In addition to these factors, CHD most often develops when cholesterol aggregates on the inner walls of the heart arteries, causing them to narrow and block blood flow. This condition is called atherosclerosis.
Medical studies have shown a clear correlation between heightened free LDL cholesterol in the body and CHD occurrence.

A study published in The New England Journal of Medicine sought to discover whether the reverse is true: Could chronically low LDL cholesterol protect people against CHD? As it turns out, the answer may be encoded in the patient’s genes.
Did you know that some rare variants can offer protection from CHD? The majority of the population does not have them, which is normal. However, the folks that do get some extra protection.
You can use the Nebula Genome Browser to search for these variants in your genome. Keep reading to learn how to do it!
The Study
In their work, the authors evaluated the Atherosclerosis Risk in Communities (ARIC) study, a longitudinal, biracial cohort designed to assess subclinical and clinical atherosclerosis. The study included four diverse communities of 4000 subjects aged 45-64 and followed the participants from 1987 to 2003.
The extensive initial examination included medical, social, and demographic data. The study recorded cardiovascular events through annual follow-ups and hospital data on deaths and discharges. They defined CHD as
- a definite or probable myocardial infarction
- a silent myocardial infarction detected by diagnostic procedures
- death due to CHD
- a coronary revascularization procedure (coronary bypass graft, coronary angioplasty, or coronary atherectomy)
The ARIC study examined the PCSK9 alleles encoding variants Y142X, C679X, and R46L. Some genetic data was lost, either due to assay failure, participants not following up, or a few participants who asked that their DNA not be used for research.
The authors analyzed the remaining data for 3363 Black subjects and 9524 White subjects.
Results
Overall, they found three PCSK9 variants associated with lower plasma LDL cholesterol levels were also protective against CHD. Two variations are mutations that cause a truncated protein (426C→G, encoding Y142X, and 2037C→A, encoding C679X), while the other mutation causes a change in the protein sequence (137G→T, encoding R46L). Population-wide, the frequency of having one of the two protein-truncating mutations was about 2% in the Black population and less than 0.1% in the White population. In addition, 81% of the Black population with one of these alleles had LDL cholesterol in the lower 50th percentile.
On the other hand, the 137G→T variant had a frequency of 3.2% in White subjects and 0.7% in Black subjects. Once again, most individuals with this mutation had lower LDL cholesterol.
The protein-truncating mutations lowered plasma levels of LDL cholesterol by about 40 mg per deciliter (1.0 mmol per liter) and were associated with an 88% reduction in the incidence of CHD. The third variant (the one that does not truncate the protein) lowered LDL cholesterol levels by about 20 mg per deciliter (0.5 mmol per liter) and was associated with a 50% reduction in CHD.
Conclusions
While there are many confounding variables when it comes to predicting CHD (lifestyle, comorbid conditions, etc.), this study attempted to understand the genetic factors that influence LDL cholesterol levels and CHD risk. It shows that even moderately lower LDL over a lifetime decrease incident CHD.
The 88% reduction in CHD events in the Black population was despite multiple other risk factors. Notably, many subjects had hypertension, almost one-third smoked, and nearly 20% had diabetes.
One of the limitations of the study is the exclusion of older individuals from the study cohort. The authors believe that the effect or protective alleles may weaken in older individuals. Additionally, while the protective variants reduced the incidence of CHD, the mortality rate was not significantly different between carriers and noncarriers in both racial groups.
Overall, PCSK9 could be an attractive target for LDL-lowering therapy.
Examine your genome!
| Did you know you can use the Nebula Genome Browser (available with Deep and Ultra Deep WGS) to check whether you have this protective variant? |
- Go to the Genome Browser. In the top left corner of the genome browser, you can find a search bar.
- The rsIDs for Y142X, C679X, and R46L are rs67608943, rs28362286, and rs11591147, respectively. Using the dbSNP database, you can find the genome coordinates in the format [chromosome number][chromosome location] are 1:55046549, 1:55063542, and 1:55039974 (GRCh38 reference genome).
- Copy-paste one of these two locations into the search bar and press enter.
- The genome browser will now zoom in on these locations.
- Activate the “Center Line” in the bar at the top to better see the location that you are looking at.
- You should see stacked, gray stripes. Those are your personal DNA sequencing reads that are aligned to a reference genome sequence (colored letters above). If your DNA sequence matches the reference, then the stripes are gray. If the sequence is somehow different from the reference, then you will see various letters and symbols in different colors.
For the variant rs67608943, you have a protective allele if you see a G (instead of the reference C allele indicated by gray stripes). Looking at the variant rs28362286, you have a protective allele if you see an A (instead of the reference C allele indicated by gray stripes). For the variant rs11591147, you have a protective allele if you see a T (instead of the reference G allele indicated by gray stripes). Remember that the protective alleles are rare and >98% of people don’t have them!
Citation
Cohen JC, Boerwinkle E, Mosley TH Jr, Hobbs HH. Sequence variations in PCSK9, low LDL, and protection against coronary heart disease. N Engl J Med. 2006 Mar 23;354(12):1264-72. doi: 10.1056/NEJMoa054013. PMID: 16554528.
