Anyone who consumes tobacco products, especially cigarette smokers, is susceptible to nicotine addiction. It reaches a point where the body’s intense craving for nicotine makes it hard for individuals to quit. Consequently, some might avoid social settings where smoking is prohibited or continue to smoke despite facing health complications due to their addiction.
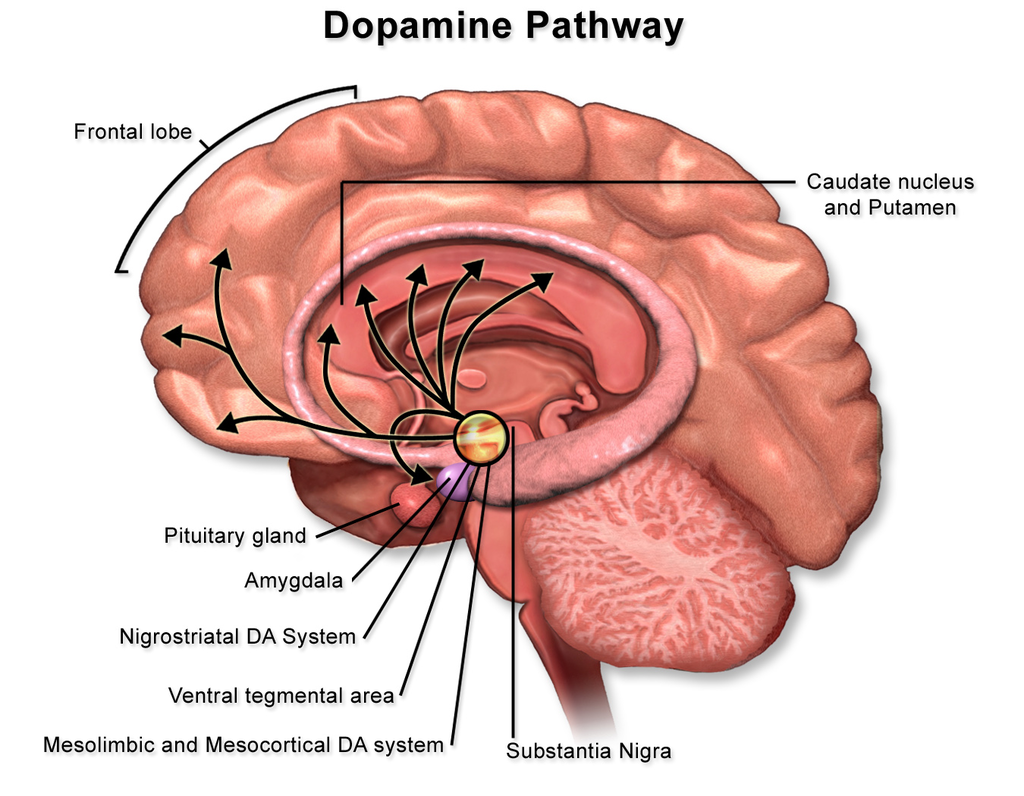
Nicotine, an essential compound in tobacco, triggers the release of neurotransmitters, notably dopamine, responsible for mood regulation. This neurotransmitter is central to the brain’s reward system. Higher dopamine levels in the brain lead to short-lived feelings of pleasure and mood enhancement. To maintain this feeling, smokers often find themselves consuming more tobacco. When they try to quit, they experience both mental and physical discomfort, symptoms of nicotine withdrawal.
The repercussions of smoking are severe, with links to fatal conditions like heart disease, lung cancer, and diabetes. While quitting smoking is challenging, many find solace in treatment plans, counseling, medical interventions, and medications.
The Study
Recent research indicates that the currently available medications for nicotine addiction, although effective, haven’t sufficiently addressed the widespread health challenges posed by tobacco consumption. The study, published in Nature Genetics, sought to identify unique genetic variants that might offer insights into genetic susceptibility to nicotine addiction and potential targets for innovative treatments.
In this study, the scientists used exome sequencing and conducted association studies related to smoking behaviors. They examined a vast cohort of up to 749,459 participants, identifying a series of variants associated with nicotine resistance.
The expansive participant count was crucial. With such a large number, the researchers were able to identify correlations with rare genetic mutations, ensuring they had a minimum of 100 carriers for each rare mutation.
Researchers categorized participants based on six smoking-related phenotypes: ever smoker, heavy smoker, former smoker, nicotine dependence, cigarettes smoked, per day (cig per day), and age started smoking.
The team then used statistical association studies to connect these phenotypes with genetic variants.
Results
The results revealed 35 rare genetic variants spread across three genes: ASXL1, DNMT3A, and CHRNB2. While the initial stages of the study also focused on the first two genes, they were eventually set aside. This decision was made based on the conclusion that mutations in ASXL1 and DNMT3A were age-acquired and related to smoking behaviors.
Significantly, the primary phenotype linked to CHRNB2 was “heavy smokers” (individuals who currently or previously smoked a minimum of ten cigarettes daily). Individuals possessing a blend of loss-of-function and missense variants in CHRNB2 had a markedly reduced propensity to be heavy smokers compared to those without these variants.

Moreover, the study identified an interplay between common and rare variants. A notable observation was that a higher genetic risk score for smoking diminished the protective effects of CHRNB2 rare variants, and these rare variants, in turn, adjusted the effect of a risk score based on common variants.
While there was an effort to pinpoint the effects of singular SNPs (single nucleotide polymorphisms), the researchers felt a more extensive sample was essential. Yet, by systematically excluding one variant at a time, a specific missense variant (rs202079239, Arg460Gly) emerged as having the most significant influence on nicotine resistance.
In essence, the study discovered that those possessing a collection of rare variants that disable CHRNB2 have a reduced likelihood of being heavy smokers. These findings could be pivotal for drug developers, opening the possibility of targeting the protein encoded by the CHRNB2 gene.
It’s vital to note that these CHRNB2 variants are uncommon. Those not carrying them have no increased smoking risk compared to the broader population. These variants simply offer added resistance!
Explore your Genome!
Gene Analysis Tool
You can use the Nebula Gene Analysis Tool (available with Deep and Ultra Deep WGS) to see whether it flags any rare variants in CHRNB2 that have a significant impact.
This tool empowers you to examine any gene in your genome and identify important genetic variants and mutations.
- When you click on the “Get Started” button your VCF file will be loaded into the Gene Analysis tool in a new tab.
- Type “CHRNB2” into the search bar at the top.
- The Gene Analysis tool will extract genetic variants in the CHRNB2 gene from your VCF file and display them to you using symbols that have different colors. The colors denote the potential importance of variants. The Gene Analysis tool determines this by referencing the ClinVar database and other resources.
- Any red and orange variants could potentially be important. Click on them to check if any of them are truncating or missense variants like the ones described in the study described above.
Genome Browser
You may also be interested in looking up the specific missense variant the study identified as contributing the most to nicotine resistance: (rs202079239, Arg460Gly).
- Go to the Genome Browser. In the top left corner of the genome browser, you can find a search bar.
- The authors associated rs202079239 with nicotine addiction resistance.
- Using the dbSNP database, you can find the genome coordinates in the format [chromosome number][chromosome location] is 1:154575801. (GRCh38 reference genome).
- Copy-paste this location into the search bar and press enter.
- The genome browser will now zoom in on this location.
- Activate the “Center Line” in the bar at the top to better see the location that you are looking at.
- You should see stacked, gray stripes. Those are your personal DNA sequencing reads that are aligned to a reference genome sequence (colored letters above). If your DNA sequence matches the reference, which is C, then the stripes are gray. If the sequence is somehow different from the reference, then you will see letters and symbols in different colors.
Citation
Rajagopal VM, Watanabe K, Mbatchou J, Ayer A, Quon P, Sharma D, Kessler MD, Praveen K, Gelfman S, Parikshak N, Otto JM, Bao S, Chim SM, Pavlopoulos E, Avbersek A, Kapoor M, Chen E, Jones MB, Leblanc M, Emberson J, Collins R, Torres J, Morales PK, Tapia-Conyer R, Alegre J, Berumen J; GHS-REGN DiscovEHR collaboration; Regeneron Genetics Center; Shuldiner AR, Balasubramanian S, Abecasis GR, Kang HM, Marchini J, Stahl EA, Jorgenson E, Sanchez R, Liedtke W, Anderson M, Cantor M, Lederer D, Baras A, Coppola G. Rare coding variants in CHRNB2 reduce the likelihood of smoking. Nat Genet. 2023 Jul;55(7):1138-1148. doi: 10.1038/s41588-023-01417-8. Epub 2023 Jun 12. PMID: 37308787; PMCID: PMC10335934.
