Cancer is an incredibly complex disease. Researchers continuously work to both understand its complexities and develop treatments. A group of scientists from Australia report the first genetic map of important genes that cause sarcoma. This is a rare cancer in muscles, bones, and other connective tissues. This type of cancer mostly derives from the embryonic mesoderm.
Published in Science, the authors identified distinct biological pathways where mutations increase the predisposition for developing sarcoma. Notably, these mutations controlled changes in telomere biology and mitotic function.
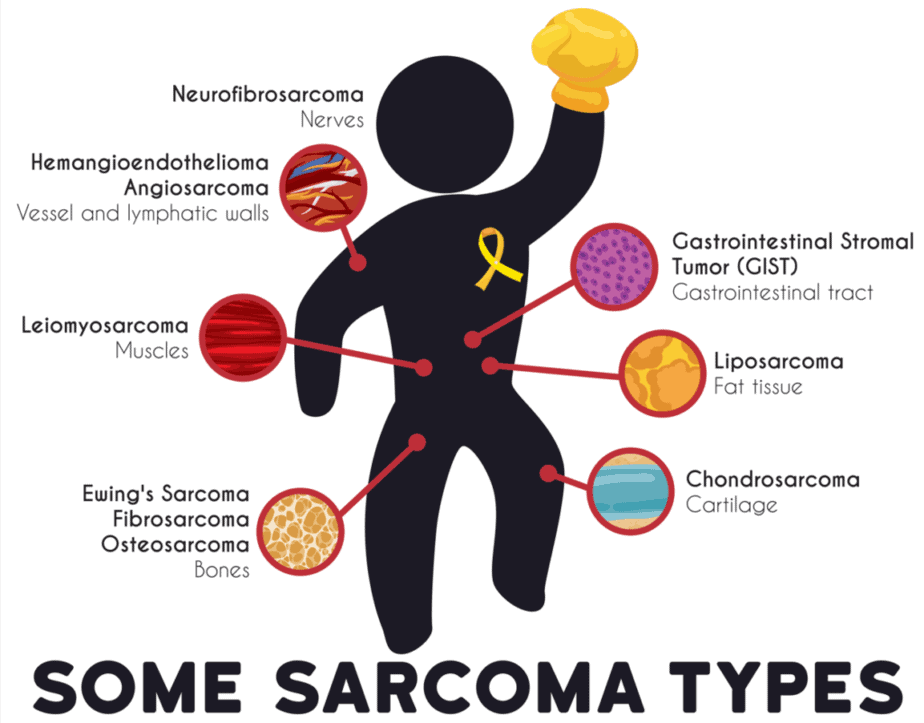
What is sarcoma?
This disease is one of the most common cancers in children and young adults. Additionally, it is often aggressive and difficult to treat. Because of its rarity, less research has been available for sarcomas than other types of cancers. This work reveals foundational knowledge of the disease so that there could be better treatments in the future.
What did they do?
Through analyzing medical history, the authors found a strong genetic basis for apparent sporadic sarcomas. Notably, the sarcomas in these individuals did not appear to occur due to radiation therapy. Futhermore, over 20% of the subjects met criteria for other known hereditary syndromes. Several had other primary cancers including breast, melanoma, second connective tissue tumors, nonmelanoma skin, prostate, colorectal, and thyroid cancers.
The authors performed whole genome germline sequencing on 1644 samples from the International Sarcoma Kindred Study (ISKS) and the Genetic Cancer Risk in the Young (RisC) studies. These samples were international and multi-institutional. Median age at first cancer diagnosis for the subjects was 47 years and 49 years at first sarcoma diagnosis.
As a control, the authors used 3205 healthy elderly controls from the Medical Genome Reference Bank (MGRB). This same research group had previously shown that this reference is depleted in cancer-associated genetic variation relative to the general population.
They used a weighted rare-variant burden test (WRVBT) to compare the variants discovered through whole genome sequencing (WGS) between the experimental data set and the control. Additionally, an ontologic analysis allowed them to focus on genes and pathways most likely to be associated with sarcoma. For example, focusing on the highest ranked genes in the WRVBT analysis and determining whether certain pathways discovered were specific for sarcoma.

Results
Overall, WGS identified over 37,000 rare single-nucleotide variants and insertions or deletions. Out of these, they discovered ~1000 known pathogenic or likely pathogenic genes while others were possibly pathogenic. Through various analytic techniques the authors determined that heritable defects in telomere and mitotic function can increase genetic predisposition to sarcoma.
Their goal was to use WGS to identify previously unknown genes and pathways that may explain sarcoma risk. Overall, they highlighted two sarcoma-specific pathways (in the shelterin and centrosome gene sets) and added 14 gene candidates to the more than 100 already known to be linked with sarcoma.
The future of cancer genomics
Studies such as this one highlight the future possibilities of WGS when looking for genetic associations in rare diseases. With publicly available datasets continuously being updated, scientists are entering an era where they can conduct stronger studies and learn more about the genetics of common and uncommon disease.
The results in this study demonstrate potential ways in which individuals develop sarcoma. Although it does not lead directly to treatment, the hope is that this type of information can be useful in early detection and eventually lay the foundation for better therapies.
Citation
Ballinger ML, Pattnaik S, Mundra PA, Zaheed M, Rath E, Priestley P, Baber J, Ray-Coquard I, Isambert N, Causeret S, van der Graaf WTA, Puri A, Duffaud F, Le Cesne A, Seddon B, Chandrasekar C, Schiffman JD, Brohl AS, James PA, Kurtz JE, Penel N, Myklebost O, Meza-Zepeda LA, Pickett H, Kansara M, Waddell N, Kondrashova O, Pearson JV, Barbour AP, Li S, Nguyen TL, Fatkin D, Graham RM, Giannoulatou E, Green MJ, Kaplan W, Ravishankar S, Copty J, Powell JE, Cuppen E, van Eijk K, Veldink J, Ahn JH, Kim JE, Randall RL, Tucker K, Judson I, Sarin R, Ludwig T, Genin E, Deleuze JF; French Exome Project Consortium; Haber M, Marshall G, Cairns MJ, Blay JY; International Sarcoma Kindred Study; Thomas DM, Tattersall M, Neuhaus S, Lewis C, Tucker K, Carey-Smith R, Wood D, Porceddu S, Dickinson I, Thorne H, James P, Ray-Coquard I, Blay JY,
Cassier P, Le Cesne A, Duffaud F, Penel N, Isambert N, Kurtz JE, Puri A, Sarin R, Ahn JH, Kim JE, Ward I, Judson I, van der Graaf W, Seddon B, Chandrasekar C, Rickar R, Hennig I, Schiffman J, Randall RL, Silvestri A, Zaratzian A, Tayao M, Walwyn K, Niedermayr E, Mang D, Clark R, Thorpe T, MacDonald J, Riddell K, Mar J, Fennelly V, Wicht A, Zielony B, Galligan E, Glavich G, Stoeckert J, Williams L, Djandjgava L, Buettner I, Osinki C, Stephens S, Rogasik M, Bouclier L, Girodet M, Charreton A, Fayet Y, Crasto S, Sandupatla B, Yoon Y, Je N, Thompson L, Fowler T, Johnson B, Petrikova G, Hambridge T, Hutchins A, Bottero D, Scanlon D, Stokes-Denson J, Génin E, Campion D, Dartigues JF, Deleuze JF, Lambert JC, Redon R, Ludwig T, Grenier-Boley B, Letort S, Lindenbaum P, Meyer V, Quenez O, Dina C, Bellenguez C, Le Clézio CC, Giemza J, Chatel S, Férec C, Le Marec H, Letenneur L, Nicolas G, Rouault K.
Heritable defects in telomere and mitotic function selectively predispose to sarcomas. Science. 2023 Jan 20;379(6629):253-260. doi: 10.1126/science.abj4784. Epub 2023 Jan 19. PMID: 36656928.
